 Finally, the number of atoms remaining at the end of this procedure is printed out along with the resulting RMS value for those atoms.ĬAUTION: Because the output RMS value and atom count are the result of a heuristic algorithm, these values should not be used for comparing relative similarity across multiple pairs of structures. back to STRAP: multiple sequence alignment also see Pymol 3D-superposition of proteins Dialog protein structure Viewer and Protein 3D-superposition with STRAP algorithms to superimpose protein 3D structures are applied to identify similarities of protein folds. In addition to plugins, some useful tools are inherently implemented in PyMOL. An initial superposition is then performed followed by up to five cycles of iterative refinement wherein atoms with per-atom deviations over two standard deviations from the mean deviation (if any) are thrown out and the fit is repeated. Load the pept.pdb PDB file of the peptide into PyMol Set the view to a wireframe rendering, by clicking S next to all in the display panel, hover over as, move to the other menu and click wire Look at the peptide. Matching side chains atoms will be included if they were provided in the selection arguments. Kastner Massachusetts Institute of Technology Can you find the backbone RMSD of two related peptides Is there a way to calculate the backbone RMSD for two different peptides that. Then, a per-atom correspondance is established between atoms in the selections. The figure below shows the three main chain torsion angles of a polypeptide. “align” first performs a per-residue global dynamic-programming sequence alignment for the input atom selections using the BLOSUM62 weightings from BLAST. 1 Secondary structure and backbone conformation 1.1 Main Chain Torsion Angles. Here, a methionine sidechain is attached to. In short, “align” is a automated multi-step superposition algorithm based on dynamic programming and iterative refinement. Peptide bonds join the carboxyl group of one segment to the amino group of the next. For each of these, InterPep2 also predicts the quality of the model as measured by DockQ-score ( Basu and Wallner, 2016a ). 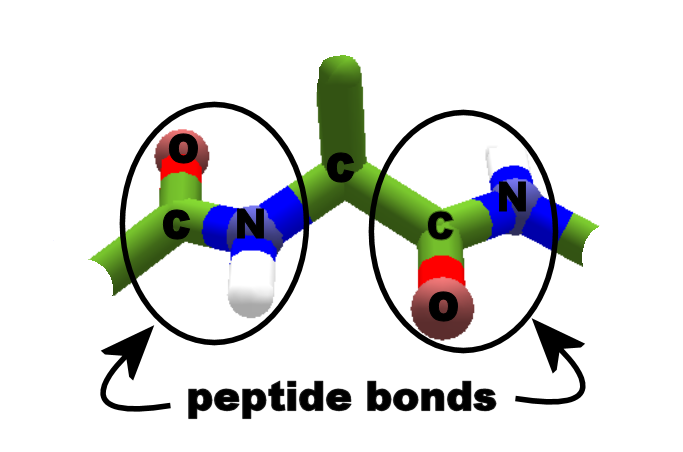
Please describe/reference the alignments algorithms used in the Pymol alignment. InterPep2 takes a protein structure and a peptide sequence as input and produces several suggested docking poses describing the peptideprotein interaction. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
